This exercise is set at the end of the first day of the Simple Shell Scripting for Scientists course. Solutions to this exercise will be examined on the next day of this course, so you must attempt this exercise before then or you may find it very difficult to follow the remaining days of this course.
This exercise was designed to be run on MCS Linux. However, it should work on most modern Linux distributions that have Python 2.7 (or any later version of Python 2), gnuplot 4.4 or higher (with PNG support), and Eye of GNOME 2.9.0 or higher (or another appropriate PNG viewer).
If you don't do this exercise on MCS Linux then you should have a look at the compatibility notes before starting the exercise.
Obtaining the files for this exercise
On MCS Linux
In order to do this exercise, you'll need to get hold of the files used in class. If you are doing this on MCS Linux, then after you have logged in, type the following:
cd "${UX}"/Lessons/Shell-Scripting-Scientists
./setup-day-one.sh
You'll be asked for a directory into which the files should be placed, the files will be unpacked to that directory and then set up for you. Look at the README file in the directory to which the files have been unpacked for instructions on how to run the script constructed in class.
On another machine
You will need to download the course files archive to your computer. You'll also need to unpack the archive and set things up with this setup-day-one.sh script. Download the archive and script to the same directory. cd to the directory into which you downloaded those two files. Make sure the setup-day-one.sh script is executable by typing:
chmod +x setup-day-one.sh
and then run it by typing:
./setup-day-one.sh
The setup-day-one.sh script will ask you for a directory into which it should unpack the archive and will then set everything up for you.
Once the archive is successfully unpacked, see the README file for details of what it contains.
The exercise
On the first day of this course we've created some shell scripts that run the zombie program for several parameter sets. These scripts make sure that the results of running zombie with a particular parameter set have names that correspond to the last parameter in the parameter set.
Change to the directory containing the zombie program and the answers and scripts subdirectories. First of all, make sure that the run-once shell script in the scripts subdirectory has been modified as we did during the course. You can do this by overwriting the copy in the scripts subdirectory with the copy provided in the answers subdirectory like this:
cp -pf answers/run-once scripts/run-once
(You only need to to do this once.)
Now type the following:
rm -f *.dat stdout-* logfile
scripts/multi-run 50 100 500 1000 3000 5000 10000 50000
This will run the zombie program 8 times, and each zombie.dat file will be renamed to something like zombie-50.dat, zombie-100.dat, etc. Now that's all very well, but it would be nice to actually see those results in graphical form.
In the gnuplot subdirectory you will find a file called zombie.gplt. This file contains some commands for the gnuplot program telling it how to turn the zombie.dat file produced by zombie into a graph. This file has been set up so that it requires a file called zombie.dat file to be in the current directory when gnuplot is started. It will create a graph from zombie.dat and store its results in the current directory as a PNG file called zombie.png.
First of all, let's copy the zombie.gplt file to the current directory:
cp -p gnuplot/zombie.gplt .
Now let us try creating a graph with gnuplot and this file. Type:
cp zombie-50.dat zombie.dat
gnuplot zombie.gplt
ls
rm -f zombie.dat
eog zombie.png &
The output of ls should show that a file called zombie.png has been created. The eog program will actually display the file for you so that you can see what it looks like. Neat, huh?
For example, the zombie.png file for the zombie.dat file that is produced by running the zombie program with the parameters 0.005 0.0175 0.01 0.01 50 looks like this:
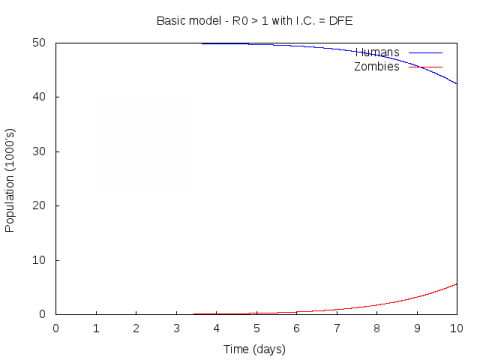
Your task is to write a shell script that will work its way through all the renamed zombie.dat files and use gnuplot (with the zombie.gplt file) to create an zombie.png file for each of them. Each zombie.png will need to be renamed or else it will be overwritten. You should rename each file to a name similar to that of the zombie.dat file from which it was generated (e.g. if zombie.png is the graph of the zombie-50.dat file, you could rename it to zombie-50.dat.png).
Hints
If you aren't sure what you should be doing, perhaps the following description of the above task will make it clearer:
We have a directory containing a number of output files, which are all called zombie-<something>.dat (where <something> is a number). What we want to do is, for each of those files, do the following:
-
Rename (or copy) the output file we want to process to
zombie.dat.mv zombie-50.dat zombie.dat -
Run
gnuplotwith thezombie.gpltfile.gnuplot zombie.gplt -
Rename (or delete if you copied the original
.datfile)zombie.dat.mv zombie.dat zombie-50.dat -
Rename
zombie.png.mv zombie.png zombie-50.dat.png
We'll be looking at your answers to this exercise on the next day of this course, so you need to have attempted it before then.
Hint: you'll probably want to use a for loop over all the .dat files in the directory.
There are two obvious ways of approaching this task: one would be to modify the run-once script we created in this course to create an zombie.png file each time it processes a parameter set. The other way would be to write a completely separate script that is designed to run after the multi-run script has completed, i.e. first run multi-run, then run this new script to create zombie.png files for all the renamed zombie.dat files previously created by run-once. For this exercise I'd like you to try the second approach, i.e. write a completely separate script. Of course, feel free to cannibalise the multi-run and run-once scripts if you find that useful.
Here are some examples of what the zombie.png files produced by your shell script for various parameter sets should look like:
-
Parameter set:
0.005 0.0175 0.01 0.01 50
-
Parameter set:
0.005 0.0175 0.01 0.01 5000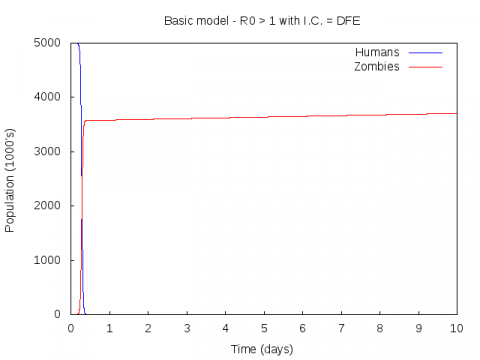
-
Parameter set:
0.005 0.0175 0.01 0.01 50000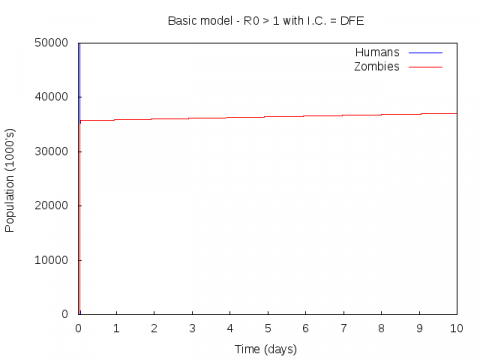
Compatibility notes
If you are not doing this exercise under MCS Linux then you should be aware of the following issues which may arise when using other Linux/Unix systems:
-
The version of the
zombieprogram provided in this archive is written in Python 2. It has only been tested with Python 2.7, although it will probably work with any version of Python 2 from Python 2.5 onwards (no promises, though). -
You may have problems if you are using a shell other than bash. You may also have problems if you are using a version of bash earlier than version 2.04. Note that the scripts used in this course were all written to run under bash 4.1, but it is believed that they will run under bash 2.04 or higher (no promises, though).
-
The
gnuplotcommands used have only been tested withgnuplot4.4. They will probably work withgnuplot4.0 or higher, but this has not been tested. -
The version of
gnuplotyou use must have PNG support. This usually depends on how your version ofgnuplothas been compiled. To find out if it has PNG support, startgnuplotand type the following:show version longIf under
Compile optionseither+PNGor+GD_PNGis listed, then your version ofgnuplothas been compiled with PNG support. -
If you do not have Eye of GNOME (
eog) on your system, almost any other PNG viewer will do. Most modern web browsers can view PNG files. A list of some applications that support the PNG format is given here.
